Dr. Vera Krymskaya is a Professor of Medicine at the University of Pennsylvania. The Krymskaya lab focuses on preclinical and translational research in rare and common lung diseases from pulmonary lymphangioleiomyomatosis (LAM) to asthma. We sat down with Dr. Krymskaya to learn more about her work and get her thoughts on the current state and future of preclinical rare lung disease research.
A: My background is in radiation biology and biophysics. During my graduate training at the Institute of Biophysics at the Russian Academy of Sciences, I studied effects of chronic ionizing radiation on plants and animals and became very interested in signal transduction. Eventually, this led me to a post-doctoral training at the Cardiovascular Institute at the Russian Academy of Medical Sciences in Moscow, Russia. Following this, I came to the United States, University of Pennsylvania, on an NIH scholarship to continue working as a post-doc studying asthma.
I was able to apply my knowledge of signal transduction and cell biology to airway remodeling in asthma. Then, as my career progressed, I felt I needed to branch out and to develop a separate project. I then learned about a rare genetic lung disease that had abnormal smooth muscle-like lesions and used LAM to study how abnormal smooth muscle grow, and now lymphangioleiomyomatosis (LAM) has been my lab’s primary focus for the past several years.
I am currently a Professor of Medicine with tenure, at the first medical school in the United States founded by Benjamin Franklin.
Online profile/Lab:
A: I began my pulmonary research in asthma, studying signal transduction and airway remodelling. I then began exploring other smooth muscle diseases and came across LAM. LAM is a rare genetic lung disease affecting almost exclusively women of childbearing age, destroying the lung and eventually leading to the loss of pulmonary function requiring lung transplantation. It is our lab’s goal and focus to identify the causes of this devastating lung destruction and provide novel treatment(s) for the disease.
A: LAM disease research provides great opportunities for a large learning curve and lots of areas to discover with relevance to common diseases. Studying rare lung disease is very important as usually it is a monogenic disease where you have dysregulation of one gene and a clear phenotype. Rare disease research is uncharted territory since there are many opportunities to make novel discoveries that can lead to treatment.
From my experience with LAM disease alone, research in my lab used Rapamycin to completely block the growth of LAM cells in 2002, and then in 2003 the first clinical trial began. It was an extremely rewarding experience how quickly you can go from basic and translational lab work to make a difference in patients’ lives.
A: Prior to my research with LAM, it was known in the scientific community that it is a genetic disease. However, it was not known which specific cells are affected by mutation of the TSC2 gene caused the disease or what cause the LAM cell growth in the lung. My lab was able to identify the function of the TSC2 gene as a negative regulator of the mTORC1 pathway. This was a very important discovery crucial to the understanding of the cause of LAM disease and begin development of in-vivo modeling.
LAM has a history of problems with animal model generation because the gene that produces LAM is involved with a very important pathway in every cell in the body. Usually, if you just knock out that gene, it is lethal. Therefore, from a genetic perspective, it is very hard to develop an in-vivo model. We were able to create a syngeneic TSC2-null mouse tumour model, published in Science Translational Medicine in 2012, which is now used by many labs. However, we still wanted to create a model where we could delete the TSC2 gene in lung-specific LAM-like cells, which would create a disease model more closely resembling the human instance of disease, which is typically the main goal of in-vivo modelling.
Recently, in our Nature Communications 2020 publication, we have been able to create a genetic model by deletion of the TSC2 gene in mouse lung cells called mesenchymal progenitor cells, leading to an age- and sex-linked structural and functional decline in the lung.
Publication references:
A: Animal models are crucial for basic and translational research. You can work with cells, which is very important. However, the next step is a transition to animal models for an in-vivo perspective to understand cellular research in a more complex in vivo model. It is important to fully understand the pathways, cell communication and signal transduction in a more complex, in-vivo, model to fully study and understand the disease, which is critical for finding novel treatments and potentially cure the disease.
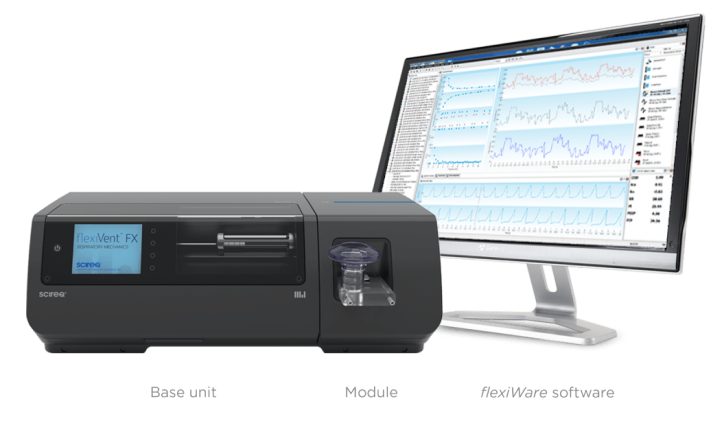

A: The flexiVent was crucial to understanding the functional impairments of the lungs in our mouse model of LAM. First, we identified the morphological changes, and subsequently, we needed to understand and quantify the actual functional impairment associated with these changes.
The flexiVent was crucial to our understanding of the functional impairment of the lung that could not be obtained via simply looking at the morphological data. In our LAM model, we saw a common characteristic of enlarged air spaces and subsequently quantified this functionality using the flexiVent.
In the recent publication using our genetic model, we studied 12 weeks and 24 weeks-old animals and were able to identify age- and sex-difference in lung function impairment. Although male models experience a change in lung morphology and structure, only females experienced the impairment of lung function. These characteristics are similar to cases seen in the human population of patients with sporadic LAM and another genetic disease TSC-associated LAM.
Additionally, we have also used the flexiVent for another rare lung disease looking at Birt-Hogg-Dube´ (BHD) syndrome, which is a rare autosomal-dominant disorder that affects the lung, skin, and kidney. In the lung, 80%–100% of patients with BHD develop multiple thin-wall cysts without evidence of neoplasia, inflammation, or fibrosis. We used the flexiVent to evaluate the functional impairment in our genetic FCLN model and impact on cyst development and alveolar space enlargement.
Publication references:
A: I think the satisfaction from seeing such advancement in our knowledge of LAM via the successful development of a challenging novel animal model, which can be very useful experimental tool to study not only rare lung disease LAM, but also other common lung disease characterized by activation of mTORC1 such as chronic obstructive pulmonary disease (COPD), pulmonary arterial hypertension (PAH), and idiopathic pulmonary fibrosis (IPF).
We have been working on developing an animal model since 2003, including a xenographic model with human tissues, an immunocompromised/immunodeficient, and genetic models. The ultimate goal, however, is to create the genetic mouse model because it is more controlled and translatable to human disease. Therefore, I am delighted that we were able to generate our current genetic model because it links specific mesenchymal cell types to the LAM disease. The LAM cell of origin is not well understood. We were able to identify this by looking at the origin of disease in the human LAM lung which occurs in the mesenchymal lineage, and then create a specific cell type knock-out of the gene in mice created our in-vivo model. It was very exciting to have a direct link between the clinical manifestation of the disease in humans and pre-clinical animal models.
This model, developed from rare-disease research with LAM, also provides many different opportunities for lung research such as ageing, sex-specific lung diseases, impairment of lung function. Dissecting different cell lineages from the lungs and analyze gene expression will help to understand cellular crosstalk with TSC2-null mesenchymal cells.
I think we’re at the beginning of a new wave of knowledge coming from single RNA cell sequencing in terms of cell lineages, their origin, gene expression and receptor-ligand crosstalk. Ten to fifteen years ago, studying the changes in genes was very novel and not well understood, but now, bioinformatic technology has caught up with us. We can take this knowledge and analyze cross-talk between cells, and it opens up a whole new world of knowledge and opportunities for discoveries of potential treatments. The idea being, we can take discoveries from novel areas like rare lung disease and expand our knowledge exponentially with the combination of collaborative research and technology.
My main advice for new researchers is to pick something you like and are most passionate about and focus on this. Then you can apply your skills and knowledge to study different angles of disease.
A: That’s a great question. We will stay with rare disease LAM, of course, as we were lucky to be part of the efforts led to FDA approval of a treatment for LAM (Rapamycin), which has extended life expectancy from seven years to about 21 years, which is terrific and feels great to have impacted lives in this way. However, we still need to understand how we can restore lung structure and lung function. This is also relevant to diseases outside of LAM, such as COPD, which also experiences lung structure degradation and dramatic reduction of lung function with age.
Studying rare diseases such as LAM helps to understand specific pathways, then apply those specific pathways to other common lung diseases such as COPD and cancer. This is our goal now, to study LAM, build up on LAM, and expand our knowledge to other lung diseases.
If you have any questions about the flexiVent for rare lung disease studies, please contact us!